viziphant.gpfa.plot_trajectories_spikeplay¶
- viziphant.gpfa.plot_trajectories_spikeplay(spiketrains, returned_data, gpfa_instance, dimensions=[0, 1], speed=0.2, orthonormalized_dimensions=True, n_trials_to_plot=20, trial_grouping_dict=None, colors='grey', plot_group_averages=False, hide_irrelevant_neurons=False, plot_args_single={'alpha': 0.4, 'linestyle': '-', 'linewidth': 0.3}, plot_args_average={'alpha': 1, 'linestyle': 'dashdot', 'linewidth': 2}, plot_args_marker_start={'label': 'start', 'marker': 'p', 'markersize': 10}, eventplot_kwargs={}, slider_kwargs={}, animation_kwargs={'blit': True, 'repeat': True}, figure_kwargs={})[source]¶
This function allows for 2D and 3D visualization of the latent space variables identified by the GPFA.
Optional visual aids are offered such as grouping the trials and color coding their traces. Changes to optics of the plot can be applied by providing respective dictionaries.
This function is an adaption of the MATLAB implementation by Byron Yu which was published with his paper: Yu et al., J Neurophysiol, 2009.
- Parameters
- returned_datanp.ndarray or dict
When the length of returned_data is one, a single np.ndarray, containing the requested data (the first entry in returned_data keys list), is returned. Otherwise, a dict of multiple np.ndarrays with the keys identical to the data names in returned_data is returned.
N-th entry of each np.ndarray is a np.ndarray of the following shape, specific to each data type, containing the corresponding data for the n-th trial:
latent_variable_orth: (#latent_vars, #bins) np.ndarray
latent_variable: (#latent_vars, #bins) np.ndarray
y: (#units, #bins) np.ndarray
Vsm: (#latent_vars, #latent_vars, #bins) np.ndarray
VsmGP: (#bins, #bins, #latent_vars) np.ndarray
Note that the num. of bins (#bins) can vary across trials, reflecting the trial durations in the given spiketrains data.
- gpfa_instanceGPFA
Instance of the GPFA() class in elephant, which was used to obtain returned_data.
- dimensionslist of int, optional
List specifying the indices of the dimensions to use for the 2D or 3D plot. Default: [0, 1]
- speedfloat, optional
The animation speed. Default: 0.2
- orthonormalized_dimensionsbool, optional
Boolean which specifies whether to plot the orthonormalized latent state space dimension corresponding to the entry ‘latent_variable_orth’ in returned data (True) or the unconstrained dimension corresponding to the entry ‘latent_variable’ (False). Beware that the unconstrained state space dimensions ‘latent_variable’ are not ordered by their explained variance. These dimensions each represent one Gaussian process timescale $tau$. On the contrary, the orthonormalized dimensions ‘latent_variable_orth’ are ordered by decreasing explained variance, allowing a similar intuitive interpretation to the dimensions obtained in a PCA. Due to the orthonormalization, these dimensions reflect mixtures of timescales. Default: True
- n_trials_to_plotint, optional
Number of single trial trajectories to plot. If zero, no single trial trajectories will be shown. Default: 20
- trial_grouping_dictdict or None, optional
Dictionary which specifies the groups of trials which belong together (e.g. due to same trial type). Each item specifies one group: its key defines the group name (which appears in the legend) and the corresponding value is a list or np.ndarray of trial IDs. Default: None
- colorsstr or list of str, optional
List of strings specifying the colors of the different trial groups. The length of this list should correspond to the number of items in trial_grouping_dict. In case a string is given, all trials will share the same color unless trial_grouping_dict is specified, in which case colors will be set automatically to correspond to individual groups. Default: ‘grey’
- plot_group_averagesbool, optional
If True, trajectories of those trials belonging together specified in the trial_grouping_dict are averaged and plotted. Default: False
- hide_irrelevant_neuronsbool, optional
If True, neural activity will be shaded according to the influence of a neuron on the chosen latent dimensions. The influence is estimated as a normalized L1-norm of the columns of the pseudo-inverse of Corth matrix:
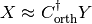
where
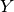 is (zero-mean) neuronal firing rates, estimated from
spikes, and
is (zero-mean) neuronal firing rates, estimated from
spikes, and 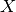 - latent variables.
Default: False
- latent variables.
Default: False- plot_args_singledict, optional
Arguments dictionary passed to ax.plot() of the single trajectories.
- plot_args_averagedict, optional
Arguments dictionary passed to ax.plot() of the average trajectories. if ax is None.
- plot_args_marker_startdict, optional
Arguments dictionary passed to ax.plot() for the marker of the average trajectory start.
- eventplot_kwargsdict, optional
Arguments dictionary passed to
plt.eventplot(). Default: {}- slider_kwargsdict, optional
Arguments dictionary for a slider passed to
ax.axvline(). Default: {}- animation_kwargsdict, optional
Arguments dictionary passed to
animation.FuncAnimation().- figure_kwargsdict, optional
Arguments dictionary passed to
plt.figure(). Default: {}
- Returns
- figmatplotlib.figure.Figure
- axesmatplotlib.axes.Axes
- spikeplaymatplotlib.animation.FuncAnimation
Matplotlib animation that can be saved in a GIF or a video file.
import matplotlib.animation as animation spikeplay.save("gpfa.gif") writergif = animation.FFMpegWriter(fps=60) spikeplay.save("gpfa.mov", writer=writergif)
